stimulus
Why Stimulus?
Better data equals better models. Refining raw bio-data is hard (it is an entire field called bioinformatics!).
Stimulus provides a systematic approach to optimise bio-data for modeling (how should I process the data? is the model gonna learn better?).
Our solution
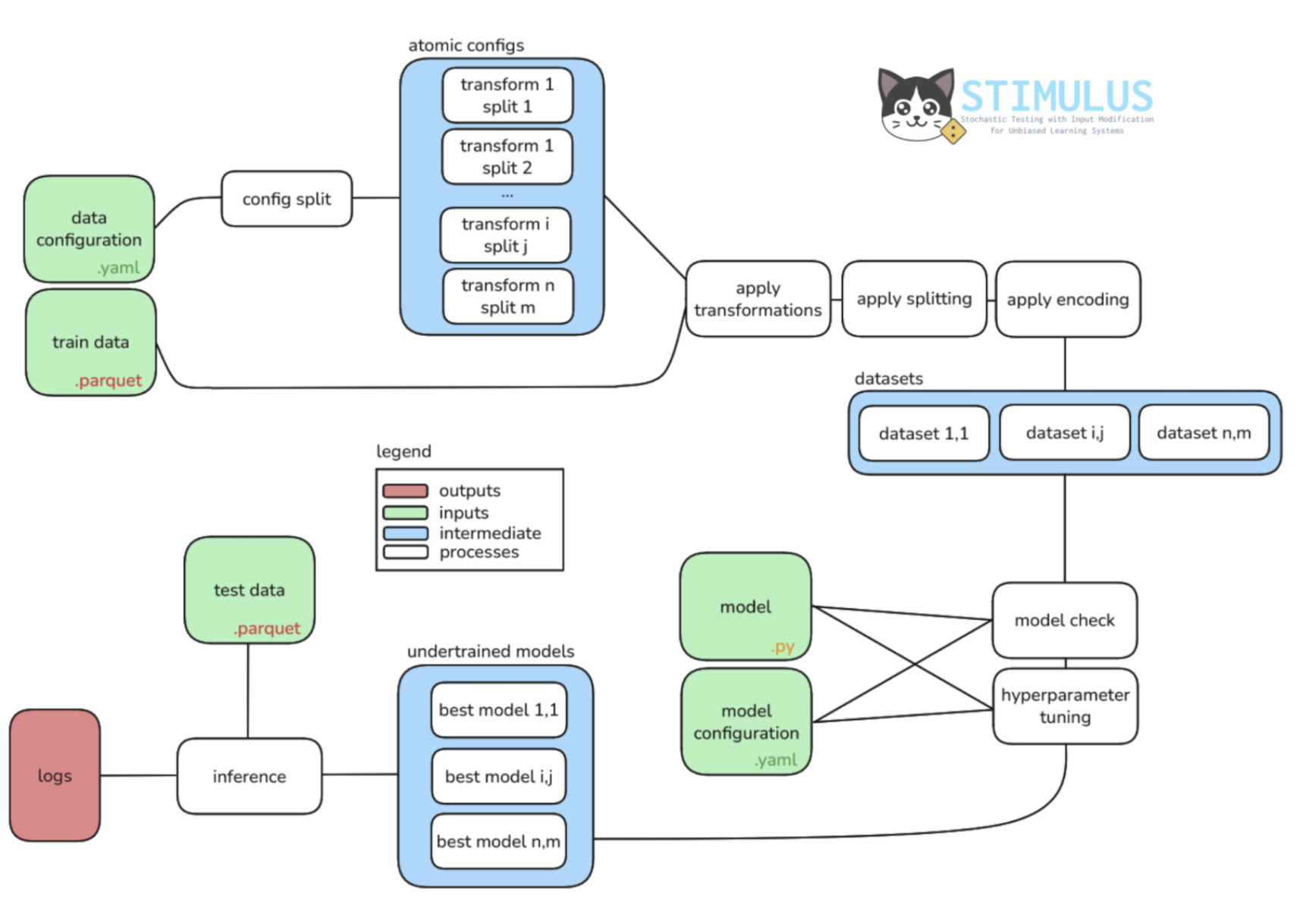
We created Stimulus, an end-to-end framework that connects data procedures (eg., bioinformatics processing, transformation, splitting, etc.), with hyperparameter optimization, training, and evaluation - all in one place.
To connect the two words (bio and DL), we used a dual implementation. The main functionalities common in DL (eg., integrating PyTorch models, calling hyperparameter tuning, configuration handling, etc) are implemented as a Python toolkit (Github: mathysgrapotte/stimulus-py). While a modular end-to-end orchestration pipeline is implemented in Nextflow and deposited in nf-core (Github: nf-core/deepmodeloptim) to not only enable systematic exploration at scale, but also connection to the bio ecosystem.